It’s only natural
An evolving series of posts on evolution … Part II
I’d suggest a fair number of people have the wrong view of natural selection. They see it as some sort of ‘theory’. It isn’t. Let’s explore that provocative assertion.
Natural Selection
The term ‘natural selection’ describes a process that will predictably occur in certain circumstances. It is something you can build and watch happening. Here are those circumstances:
A mechanism exists for making copies of things;
These copies are imperfect enough for the differences among them to matter;
There is a Grim Reaper that deletes things preferentially, based on one or more criteria that touch on those differences.
It would seem that the circumstances I’ve stated contain a lot of weasel words. For example, what sort of ‘things’ are we talking about? What does ‘imperfect enough’ mean, and how does the Grim Reaper choose, precisely? But it turns out that the ‘things’ can be pretty much anything, and the preferential choice among the imperfect copies can be pretty general too. The differences don’t even need to be that large, especially if we run this copy-and-delete mechanism for a while.
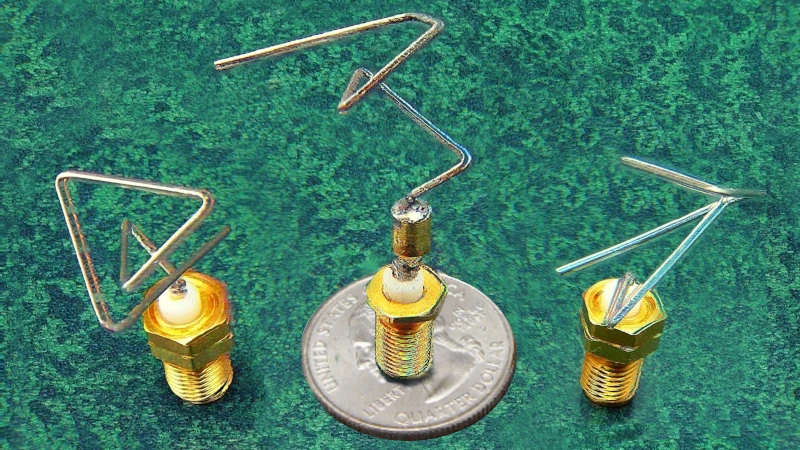
Take the picture at the start of my post. These ugly pieces of bent wire are actually rather special, especially the one in the middle, resting on a US quarter. ST5 was a NASA space mission that tested ten new technologies. These wedding-cake-sized satellites were launched on 22 March 2006, testing miniature digital communication tech.
The design of the antenna tested and used on these satellites proved ‘challenging’, as they had a whole lot of very specific requirements. The smart chaps at NASA decided to test that ugly little antenna against the best human-conceived designs (‘quadrifilar helical antennae’). But how did they come up with new antennae? Natural selection: they created a little “antenna constructing language” and had it build multitudes of small antenna models, starting with a basic design. They then automatically evaluated each model against the many criteria it needed to meet, weeding out the less fit variants.1
That weirdly bent bit of wire did the job.
Evolutionary algorithms v biology
And of course—provided you have a suitable way to test your model—you don’t need to build each antenna. Everything can be done using software alone. There are limitations: the computer power required for testing can be quite spectacular, depending on the variety of criteria being evaluated, and the building blocks that are available.2
With biological organisms, we have it a lot easier than people playing with evolutionary algorithms on a computer. We have:
An impressive set of building blocks.
A very effective Grim Reaper: Mother Nature is a merciless bitch.
Prolific mechanisms for making imperfect copies of things at a molecular level.
A stupendously huge testing space, deployed in deep time.
Here too, we need to get some things clear. In terms of the sheer number of ‘things happening in a second’, a supercomputer is quite pathetic in comparison to—well—a cup of soup. One of our current crop of supercomputers from the past few years can manage perhaps a few exaflops, that is over 1018 floating point operations per second.
A billion billion multiplications. That sounds like a lot, but the flow of information is very specifically directed, and the room for “lateral interaction” of information is quite limited. In contrast, your average cup of soup (say a metric cup of 250 ml) contains about 14 moles of water, or about 8×1024 molecules, rearranging themselves continuously, and interacting almost willy-nilly. Hydrogen bonds form and dissociate within about 2 picoseconds. This means a mind-blowing 1036 interactions per second.
We can extend this outrageous comparison. It may take the entire resources of a supercomputer deployed for simply ages to reliably model a single protein molecule in that cup of soup. We have a whole cupful, all interacting, forming and twisting and reforming in a seething welter of interactions.
And even when it comes to ginormous molecules like DNA, there is an astonishing capacity for things to happen. Please contemplate the following video of DNA replication:
Nucleic acids—a quick review
A normal human cell contains about 6 billion base pairs of DNA.3 If you were to unroll this, it would stretch for about 2 metres, as there’s about 0.34 nanometres between base pairs. Some estimates put the number of cells in the human body at about 30 trillion, so end-to end we’re looking at 60×109 km of DNA, or 64 times the length of the Earth’s orbit. I’ve seen estimates that the amount of DNA made in a single human body during their lifetime would extend for about a light year!
When copying DNA in such large amounts, a lot can go wrong. Fortunately, we have amazing cellular machinery that defends us. Everyone should know that the genetic code in DNA has simple building blocks: just four of them. These 4 base pairs naturally pair up: adenine (A) with thymine (T), and guanine (G) with cytosine (C).
In theory, DNA is therefore easy to copy: separate the strands, and pair each base with its complement. But DNA replication happens at a ferocious rate (as you can see in the video) and sometimes there’s a misfit. Fortunately, we have DNA proofreading mechanisms that are remarkably good at picking this up. The bad base is cut out so efficiently that replication errors for DNA are literally one-in-billion. The analogy here is if you typed continuously at 60 words per minute for six years, you’d make one error.
When we look at viruses that lack proofreading when they make copies, things are very different. They are often a thousandfold less accurate, which provides us with the opportunity to study natural selection pretty much in real time.
For example, human immunodeficiency virus is notorious for rapidly developing resistance against antiviral drugs. Once we’d worked out the cause of AIDS, we started exploring “anti-retroviral” drugs. The problem was that mutation rate. Most mutations are either neutral or harm the virus, but natural selection steps in.
The Grim Reaper is unforgiving here, in both senses. Those virions that are even a bit resistant to an anti-retroviral drug get an advantage—and with every iteration, this has the potential to improve. When you’re infected with HIV, you form billions of new virions every day, so the virus has a lot of room to play. Resistance to effective agents develops with surprising rapidity.
Understanding this rapid turnover allowed Perelson and colleagues to create mathematical models of virus dynamics. Mathematically, combination therapies could be shown to work. Highly-active anti-retroviral therapy had arrived: HAART. Now we have highly effective ‘ART’ options that have transformed HIV infection from a death sentence to a manageable, chronic condition. We can defeat the powerful natural selection afforded to the virus by its lack of fidelity in making copies of itself.4
The eyes have it
I ended off my last evolutionary post⌘ by noting that the ‘seven spanning’ G-protein coupled receptors (GPRCs) we explored have a wealth of uses in humans. One of these is for vision. We found that the box jellyfish also uses similar 7TM receptors to see.
You’d think that with all the potential machinery that might arise over deep time, there would be a near-infinite variety of proteins, and a near-infinite variety of organisms. And to a certain extent, Nature explores every niche. But the remarkable thing here is that we see recurring themes. We see this at the tiniest level, and we see this with the end-product of all those genes and proteins doing their thing—the actual organism.
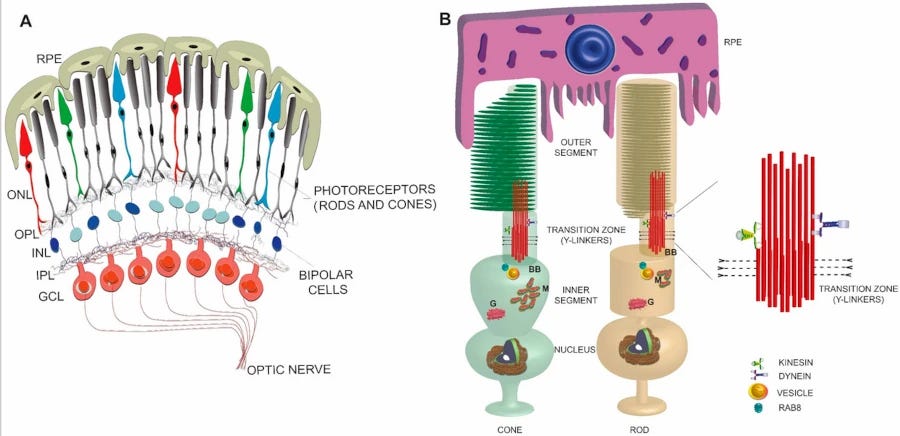
Back in 1876, Franz Boll noticed that a rose-coloured pigment in the retina of the eye bleached on exposure to light. This acquired the name rhodopsin.5 It’s found in the rod cells, which react to low light. Like most visual pigments, rhodopsin is bound to vitamin A, here a variant confusingly called retinal. Retinal retinal.
It turns out that vitamin A is particularly useful for detecting light, whether you’re a vertebrate, an insect or a squid. Bound to proteins called opsins, retinal changes its shape when a photon of the right wavelength strikes it. This in turn tweaks the shape of the opsin, and a signal cascade is kicked off. We have vision!
We’ve now discovered light-sensitive proteins called bacteriorhodopsins. Let’s call them ‘BRs’ for short. Remarkably, these are seven-segment membrane spanning proteins that are bound to retinal, and are light responsive. They have similar protein structure, colour sensitivity and light reactions to rhodopsin in organisms with nuclei (Eukaryotes).
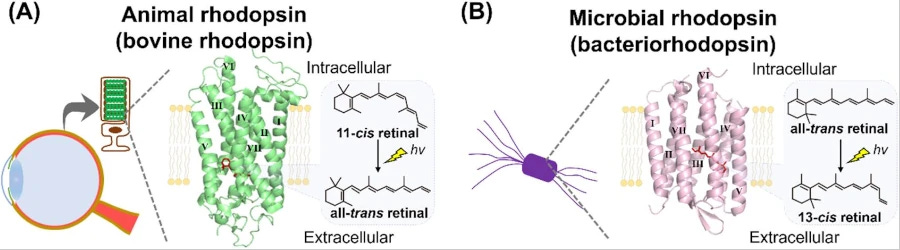
Convergent evolution
But that’s where the similarity stops. There is no associated G protein. The salt-loving Archaea that use BRs most enthusiastically seem less interested in detecting light: instead, they harness the energy of green light, use it to create a pH gradient, and then turn this into high-energy phosphate bonds to power their cells.6
Recent analysis suggests that despite their remarkable similarities, BRs and animal rhodopsins share no common ancestor. We’ve discovered many thousands of BR-like proteins, and analysed them in detail. BR-like proteins are found not only in Archaea, but also in fungi, algae, bacteria and even viruses. They do all sorts of things, but these mostly relate to pumping various types of ion, often controlled by light exposure (‘light gating’). There seem to be just too many dissimilarities for these ion pumps to be related to opsins.
There is a host of similar examples of convergent evolution at the molecular level. We find this with many enzymes; there’s also the glorious observation of how the protein prestin has acquired similar features in dolphins and bats, driven by natural selection. There’s no ultrasound-detecting ancestor common to dolphins and bats.
These patterns extend all the way up from molecules to the whole animal. Previously we explored how the finest details leak upwards into the ‘big picture’.⌘ Nowhere is this more tangible than when we move up from proteins to whole organisms. Different groups may converge towards remarkable similarity. The classical example here is flight: the wings of bats, birds and pterosaurs are remarkably similar in many ways, despite these respectively being mammals, dinosaurs, and reptiles. Another is how the hydrostatic (inflatable) penis has been discovered half a dozen times by different groups of animals. A third example is carcinisation: crustaceans that have nothing to do with crabs seem to be drawn by natural selection into looking like crabs.
Conversely some organisms only seem unrelated. The heady mix of natural selection and random variation over time has obscured the relationship. It’s rather fun to pair up whales and hippos, or elephants and hyraxes, and a bat is far closer to a horse than a rat!
Molecular clocks
Teasing out this sort of thing is difficult. There are opposing forces. We clearly have functional requirements imposed by the Grim Reaper. In our 7TM example, certain functions seem to be favoured by having precisely seven trans-membrane protein segments. Six may be too simple; eight or nine, unnecessarily complex.
It’s also clear that if some proteins are to work, certain parts are critical. If your opsin can’t bind retinal, it’s a non-starter. You can work out that some motifs (patterns or themes) will be highly conserved. As we’ve seen, some may even converge from different origins.
But what about boring changes? Entropy and random variation are powerful forces too. First, the genetic code is redundant: we have 64 codes and far fewer actual amino acids. Over time, a mutation can occur where a different triplet of bases codes for the same amino acid.
Also, let’s say a mutation occurs at a site that isn’t critical for binding something or ensuring that a protein bends in a specific way. For example, amino acids in those helical parts of proteins that span cell membranes tend to be water-avoidant (hydrophobic). Switch one hydrophobic amino acid with another, and very little may change.
Back in 1962, Linus Pauling and Émile Zuckerkandl rather empirically noticed that if you compare haemoglobin from different species, the amino acid sequences diverge more or less according to how divergent the species themselves are, based on other evidence like the fossil record. It’s as if the random mutation of codes for non-critical amino acids ticks like a clock.
You can also deduce that there are going to be dozens of things that influence how fast this clock ticks. Species that reproduce faster will have more opportunity to vary, compared to those that live longer and breed infrequently. Albatrosses seem to have slow clocks, for example. The clock may appear to tick faster if you measure ‘variation’ that simply reflects short term changes that haven’t ‘bedded down’ in the genome. Once a mutation at a specific point has occurred, it may be replaced by another mutation at that same point, and from then on you’ll see just one mutation rather than two, making the clock seem slower. It’s all a bit messy.
In addition, the calibration mechanisms you use may be off—are you sure that the fossil you’re calibrating against is representative, and at what point did two species (or genera, or families or orders) branch off?
This digression introduces two concepts. One is that, if we are to put molecular clocks to good use, we must acknowledge their limitations, use all the evidence we have, and join things up using good Science.⌘7 No theory can be cast in stone.
The second pretty obvious concept is that if we’re going to use molecular clocks to answer the question “When did these species separate?”, we need to be clear about what we mean by a ‘species’. And this introduces a lot of secondary questions. How do we tell two species apart? How do species even come about? We’ll confront all of these in my next post. It’s only natural that we also get to meet our fabulous friend, Charles Darwin.
My 2c, Dr Jo.
⌘ This symbol is used to indicate posts where I’ve discussed the flagged topic in more detail.
The image at the start is a composite I made from two images, here (NASA) and here.
Their initial success ST5-3-10, looked different from ST5-33-142-7 shown, which represents further evolution based on changing requirements.
The ideas behind ‘genetic algorithms’ go back to Alan Turing in 1948. John R Koza, who also invented the scratch card, tirelessly promoted them. They have produced competitive results in a vast number of areas that include design of electric circuits and of lenses, quantum computing, photonic crystals and solving differential equations. You can build your own genetic algorithms in Python.
Apart from red blood cells, which lack nuclei.
Strangely enough, the body’s own defences contribute to the high mutation rate. Many animals have defence mechanisms against single-stranded DNA copies of viruses. Generally, these alter the bases to make them read like nonsense. One such defence is the primate enzyme APOBEC3G, one of the ‘A3’ family of proteins that convert C to U. Unfortunately for us, HIV has a peptide that interferes with this enzyme—just enough that the enzyme increases the mutation rate substantially, but not enough to render the virus inactive.
From the Greek ῥόδον, a rose. English speakers used to refer to it as ‘visual purple’. In his long paper from 1877, Boll refers to the pigment as ‘Sehrot’, or ‘visual red’, but unfortunately Wilhelm Kühne climbed onto the bandwagon in 1878, calling it ‘Sehpurpur’. It seems he was referring to Tyrian purple, which is often pinkish-red, but this was naively translated as ‘purple’.
The quantum efficiency of this process is over 60%.
Modern methods for using all of the information are now pretty impressive. These are often Bayesian. The software can be extremely complex; the arguments about the merits and demerits of the software, even more so.


Back in the late 80s the team I was in was evolving 2D aerofoil designs. The process discovered the divergent trailing edge, but McDonnell Douglas had already patented it.
It's a very powerful tool. By the mid 90s thanks to Moore's Law and a bit of ingenuity we were applying it to whole wings including optimisation of weight and fuel volume.
Recently, I had a requirement to simulate the processes going on in a cup of soup.
I thought of making a new language "soup_lang", to simulate these interactions, to run in parallel.
It turns out, I already had the necessary hardware in my kitchen, and used water, chicken, and a few other things to introduce the necessary entropy into the calculations.
The programs (I'll call them "recipes") were not hard to write, pretty tolerant of the supplied parameters.
No Qbits were inconvenienced.